Cell, Vol. 114, 135–146, July 11, 2003, Copyright 2003 by Cell Press
Structure of the Rho Transcription Terminator:
Mechanism of mRNA Recognition
and Helicase Loading
this process by a number of factors, such as NusG,
which serve as antiterminators to ameliorate the termi-
nation properties of the enzyme (Sullivan and Gottes-
man, 1992).
Tethering of Rho to a rut site is independent of ATP
Emmanuel Skordalakes and James M. Berger*
Department of Molecular and Cell Biology
University of California, Berkeley
239 Hildebrand Hall, #3206
Berkeley, California 94720
binding and/or hydrolysis (Galluppi and Richardson,
1980; McSwiggen et al., 1988) and is mediated by an
N-terminal domain in each protomer of the Rho hexamer
Summary
(Martinez et al., 1996a, 1996b; Allison et al., 1998; Bri-
ercheck et al., 1998). Previous high-resolution NMR and
In bacteria, one of the major transcriptional termina-
X-ray crystal structures of the N-terminal mRNA binding
tion mechanisms requires a RNA/DNA helicase known
domain of Rho have provided a structural view of this
as the Rho factor. We have determined two structures
fold and its interactions with target substrates (Allison
of Rho complexed with nucleic acid recognition site
et al., 1998; Briercheck et al., 1998; Bogden et al., 1999).
mimics in both free and nucleotide bound states to
The domain consists of a three-helix bundle that rests
3.0 A
˚
resolution. Both structures show that Rho forms
on top of a five-stranded  barrel. The barrel belongs
a hexameric ring in which two RNA binding sites—a
to the oligonucleotide/oligossacharide binding (OB) pro-
primary one responsible for target mRNA recognition
tein superfamily and is primarily responsible for Rho’s
and a secondary one required for mRNA translocation
preferential interaction with cytosine-rich nucleic acids
and unwinding—point toward the center of the ring.
(McSwiggen et al., 1988; Geiselmann et al., 1992; Rich-
Rather than forming a closed ring, the Rho hexamer
ardson and Richardson, 1992; Modrak and Richardson,
is split open, resembling a “lock washer” in its global
1994; Platt, 1994). Curiously, the domain does not spe-
architecture. The distance between subunits at the
cifically recognize the 2⬘-hydroxyl of RNA and can asso-
opening is sufficiently wide (12 A
˚
) to accommodate
ciate with ssDNA in vitro, a property that has been used
single-stranded RNA. This open configuration most
to great effect for dissecting Rho function (Oda and
likely resembles a state poised to load onto mRNA
Takanami, 1972; Richardson, 1982; Brennan et al., 1987).
and suggests how related ring-shaped enzymes may
The C-terminal region of Rho contains signature se-
be breached to bind nucleic acids.
quence motifs that are hallmarks of Walker-type ATP
binding proteins. The closest homolog to Rho is the F
1
Introduction
ATP synthase, an ␣
3

3
heterohexameric complex that
assembles into rings and generates ATP from a proton-
Appropriate termination of transcription is essential for
motive force (Boyer, 1997). Consistent with this relation-
proper regulation of gene expression. In bacteria, mRNA
ship, EM studies have shown that Rho also forms hex-
transcription termination is carried out by two distinct
americ rings, although in the absence of RNA, these
mechanisms (Richardson and Greenblatt, 1996). In in-
rings exist in both closed and “notched” open forms
trinsic termination, an mRNA stem loop structure is
(Oda and Takanami, 1972; Gogol et al., 1991; Yu et al.,
thought to induce RNA polymerase to pause, promoting
2000). Notched rings have been interpreted either to be
the release of both the polymerase and the RNA tran-
missing a subunit or to be a natural functional state of
script from the template DNA. This mechanism accounts
the protein for loading RNA into the hole of the hexamer
for approximately half of the termination sites in Esche-
(Yu et al., 2000). Open Rho rings can be converted into
richia coli, and resembles the transcription termination
the closed form by the binding of extended single-
mechanism used by eukaryotic pol III. In contrast, the
stranded nucleic acids, which have been proposed to
second mechanism of transcription termination is pro-
wrap around the perimeter of the hexamer (Galluppi
tein mediated and relies on an enzyme found throughout
and Richardson, 1980; McSwiggen et al., 1988; Gan and
bacteria known as the Rho factor (Brown et al., 1981;
Richardson, 1999; Yu et al., 2000).
Opperman and Richardson, 1994).
As Rho loads onto nascent mRNAs, the single-
Since its discovery in 1969 by Roberts (Roberts, 1969),
stranded substrate is thought to bind in the hole of the
Rho has become a paradigm for understanding how
hexameric ring, where it interacts with a secondary RNA
exogenous proteins can regulate the termination of tran-
binding site in the C-terminal domains (Burgess and
scription by RNA polymerase. In vivo, Rho loads onto
Richardson, 2001a; Burgess and Richardson, 2001b).
mRNAs at a cytosine-rich region of 40 or more bases
Two regions, the Q and R loops, have been implicated
known as a Rho utilization (rut) site (Morgan et al., 1985;
in RNA associations at the secondary site (Miwa et al.,
Alifano et al., 1991). Once bound, Rho acts as a hexa-
1995; Burgess and Richardson, 2001a; Wei and Richard-
meric, 5⬘→3⬘ ATP-dependent helicase to translocate to
son, 2001b), and RNA binding in the hole of the hexa-
the site of transcription and disengage the polymerase
meric ring can stimulate ring closure when the second-
(Oda and Takanami, 1972; Lowery-Goldhammer and
ary sites become affixed to the message (Gogol et al.,
Richardson, 1974; Morgan et al., 1983; Brennan et al.,
1991; Yu et al., 2000). After loading appropriately, the
1987; Richardson, 2002). Rho can be regulated during
C-terminal domains are thought to engage in cycles of
ATP binding and hydrolysis, translocating Rho along the
mRNA until it reaches RNA polymerase and unwinds
*Correspondence: [email protected]
Cell
136
the RNA/DNA heteroduplex at the site of transcription ssDNA and another bound to ssRNA and AMPPNP. The
ssDNA-Rho complex was phased to intermediate reso-
(Lowery-Goldhammer and Richardson, 1974; Brennan
lution (6.0 A
˚
) by single isomorphous replacement from
et al., 1987, 1990; Walstrom et al., 1997; Burgess and
a tantalum bromide derivative, followed by high-resolu-
Richardson, 2001a; Kim and Patel, 2001; Stitt, 2001).
tion phasing using selenomethionine-based MAD. The
Two models have been advanced to describe the physi-
ssRNA/AMPPNP assembly was solved by molecular re-
cal nature of the translocation/unwinding reactions
placement. Both structures were refined to 3.0 A
˚
resolu-
(Richardson, 2002; Delagoutte and von Hippel, 2003). In
tion (Tables 1 and 2).
one, known as the tethered tracking mechanism, Rho
The crystalline asymmetric unit contains six Rho pro-
is thought to remain attached to the rut site while translo-
tomers assembled into a discrete hexameric particle.
cating toward the 3⬘ end of the mRNA (Steinmetz and
Each subunit is peanut-shaped, with dimensions of 65 A
˚
Platt, 1994). In the other model, Rho is thought to detach
in height, 50 A
˚
in depth, and 35 A
˚
in width (Figure 1A),
from the rut site upon the onset of translocation (Geisel-
and is composed of an N- and C-terminal domain joined
mann et al., 1993; von Hippel and Delagoutte, 2001).
together by an extended 30 residue linker. Although
Despite extensive efforts, several questions regarding
there is clear electron density for five out of six
the mechanism of Rho remain unanswered. For exam-
N-terminal domains in the hexamer, the N-terminal do-
ple, it is not known how the primary and secondary RNA
main of protomer B is partially disordered, and electron
binding sites are oriented with respect to each other,
density can only be seen for its C-terminal half.
and whether this arrangement can accommodate the
The Rho N-terminal domain consists of two subdo-
simultaneous RNA binding events predicted by the teth-
mains: a three-helix bundle followed by a five-stranded
ered tracking model. It has also remained unclear how
 barrel (Figure 1A). The  barrel is an OB-type fold,
an extended nucleic acid segment gains entry into the
which is found in a wide variety of single-stranded nu-
topologically closed-ring system of the Rho particle, a
cleic acid binding molecules that include proteins such
problem that exists for a variety of hexameric helicases.
as type II aminoacyl tRNA synthases and cold shock
To begin to address these questions, we determined
proteins (Cavarelli et al., 1993; Newkirk et al., 1994;
the structure of the full-length Escherichia coli Rho pro-
Schindelin et al., 1994). An indented face on one side
tein bound to an ssDNA recognition site mimic to 3.0 A
˚
of this subdomain forms the primary RNA binding site
resolution. The same structure bound to ssRNA and the
of Rho (Modrak and Richardson, 1994; Briercheck et al.,
ATP analog AMPPNP (adenosine 5⬘-(,␥-imido)triphos-
1998; Bogden et al., 1999). Structural comparisons of
phate) was also determined to 3.0 A
˚
resolution. Both
either the helical-bundle or the OB-fold subdomains be-
structures show that Rho forms hexameric rings in which
tween different protomers and isolated Rho N-terminal
the primary RNA binding sites of the N-terminal domains
domains solved previously by NMR and X-ray crystallog-
face toward the interior of the particle, positioning the
raphy show that the individual structures of these seg-
3⬘ end of nucleic acid exiting from the rut-recognition
ments do not vary greatly (C␣ RMSD 0.73A
˚
) (Allison et
surface directly toward the hole of the hexamer. Nucleo-
al., 1998; Briercheck et al., 1998; Bogden et al., 1999).
tide binds in a crevice between the C-terminal domains
However, when comparing both subdomains in unison,
of neighboring protomers and is liganded by residues
the overall RMSDs for the entire N-terminal domain be-
of the Walker-A and -B motifs. Secondary RNA binding
come higher (C␣ RMSD 1.1A
˚
) due to small variations in
motifs, which comprise the acceptor sites for translocat-
the relative orientations between the helical bundle and
ing along RNA, line the perimeter of the interior hole:
the OB fold. In the Rho hexamer, these differences are
six Q loops together form a narrow constriction in the
most prominent for protomers A and F.
center of the ring, while each R loop lies above the
Each of the six C-terminal domains of Rho consists of
P loop of the Walker-A motif. Strikingly, the Rho ring is
seven parallel  strands (6–13) sandwiched between
split open in both the AMPPNP bound and unbound
several ␣ helices (Figure 1A). There is clear electron
states, adopting an architecture akin to that of a lock
density for residues 155–417 in all six C-terminal do-
washer. Taken together, these structural features illumi-
mains, although the last 15 residues exhibit higher than
nate the mechanism by which Rho associates with rut
average temperature factors than the rest of the model.
sites and loads the 3⬘ end of the mRNA into the interior
The average C␣ RMSD between each of the six mono-
of the ring prior to the start of translocation.
mers in the hexamer is 0.58 A
˚
. As expected from se-
quence analysis, the fold of the C-terminal domain be-
Results and Discussion
longs to the RecA superfamily of ATP binding proteins.
A structural survey of such folds in the database shows
The Rho Monomer Structure
that this region of Rho is most similar to the mitochon-
The full-length Rho protein used for these studies is
drial F
1
ATPase (C␣ RMSD 2.1 A
˚
) (Abrahams et al., 1994;
hexameric in solution, as judged by both gel filtration
Bird et al., 1998b).
and dynamic light scattering (data not shown). Although
There are several key signature sequence motifs lo-
preliminary crystals of the purified protein grew readily,
cated in the Rho C-terminal domain. One is the P loop,
extensive optimization of initial crystallization condi-
which is part of the Walker-A motif found in all RecA-
tions was required to obtain crystals that diffracted to
family ATPases and is required for ATP binding and
a useful resolution. The key elements in this process
hydrolysis. Each P loop is formed by the 6/␣7 connec-
proved to be cocrystallization of the protein with mild
tor segment and is juxtaposed with an adjacent pro-
detergents and the use of an appropriate length and
tomer. A second important motif in Rho’s C-terminal
sequence of nucleic acid substrate. In all, two different
domain is the R loop, which connects the secondary
structural elements 11/␣12 and is located just belowRho structures were obtained: one in complex with
Rho Termination Factor Structure
137
Table 1. Data Collection and Phasing Statistics
Rho-DNA Rho-RNA
Complex Complex Ta
6
Br
12
2
⫹
Se-1 Se-2
Data set ( A
˚
) 1.245 1.0 0.9794
Resolution (A
˚
) 50–3.0 50–3.0 50–4.6 50–3.6 50–3.6
Completeness (%) 97.9 (97.8) 96.1 (95.0) 95.7 (99.1) 94.3 (94.8) 96.2 (96.7)
Redundancy 2.2 (2.2) 2.2 (2.2) 2.5 (2.5) 2.3 (2.3) 2.3 (2.3)
R
sym
(%)
a
4.6 (27.2) 5.0 (33.2) 6.9 (30.4) 6.3 (24.5) 6.7 (29.4)
I/ 15.5 (3.5) 15.3 (2.5) 10.0 (3.0) 12.9 (5.0) 11 (4.5)
Phasing Analysis Ta
6
Br
12
2
⫹
Se
Resolution (A
˚
) 20–6.0 20–3.6
R
cullis
(%,acent/cent, iso)
b
0.78/0.68 0.43/0.40
R
cullis
(%, anom)
b
0.80 0.78
Number of sites 6 83
Mean figure of merit (FOM) 0.432 0.308
Values in parentheses are for the highest-resolution bin.
a
R
sym
⫽⌺⌺
j
|I
j
– 具I典|/⌺具I典, where I
j
is the recorded intensity of the reflection j and 具I典 is the mean recorded intensity over multiple recordings.
b
R
Cullis
⫽⌺⌺||F
PH
⫾ F
P
| ⫺ F
H
|/⌺|F
PH
⫾ F
P
|, where F
P
and F
PH
are the structure amplitudes of the parent and the heavy-atom derivative and F
H
is
the calculated heavy-atom structure factor.
the P loop. A third motif, the Q loop, is formed by an full-length, ring-shaped ATPases observed crystallo-
graphically to date have been imaged with their rings8 residue segment that projects toward the center of
the hexamer. Together, the Q and R loops are thought closed. Open configurations of oligomeric ATPases
have been observed previously for only a few proteins,to form Rho’s secondary RNA binding site (Burgess and
Richardson, 2001a; Wei and Richardson, 2001a, 2001b; including the recombination protein RecA and a trun-
cated form of the bacteriophage T7 gp4 primase/heli-Xu et al., 2002).
case (Story et al., 1992; Sawaya et al., 1999). However,
the particles in these structures do not exist as discreteThe Rho Assembly
The six subunits of Rho pack laterally into a hexameric hexamers and instead assemble into continuous fila-
ments with 6
1
symmetry. EM analyses have shown thatring (Figures 2A and 2B). Subunit-subunit interactions
occur between both the N- and C-terminal domains of Rho hexamers are able to individually form both open
and closed rings (Oda and Takanami, 1972; Gogol eteach protomer. Contacts between adjacent N-terminal
domains are generated between the ␣2/␣3 connector al., 1991; Yu et al., 2000), although the purpose, function,
and oligomeric organization of the open conformationloop in the helical bundle of one protomer and the ex-
tended linker that joins the N- and C-terminal domains have remained somewhat enigmatic. Our ability to trap
and image an open conformation reminiscent of thatof the adjacent subunit. In the C-terminal domain, ␣11
in one protomer packs against the 7/␣8 and 8/␣9 seen previously by EM now suggests that this conforma-
tion is a stable and natural form accessed by the protein.junctions of the neighboring subunit. The connector loop
for secondary elements 11/␣12 is also located at the The diameter of the Rho ring is 120 A
˚
, with a large
interior hole that varies in width from 20–35 A
˚
. The widestC-terminal subunit-subunit interface and is positioned
adjacent to and above the P loop of the ATP binding point of the hole is defined by the perimeter of the
N-terminal domains, while the most narrow constrictionsite.
Strikingly, the Rho ring is split open, giving rise to a is formed by the Q loop signature sequence motifs. The
lock-washer arrangement of the hexamer arises fromglobal structure reminiscent of a lock washer (Figures 2A
and 2B). This conformation is unusual, because nearly all an upward rotation of one protomer with respect to
Table 2. Model Refinement
Rho-DNA Complex Rho-RNA Complex
Resolution (A
˚
) 20–3.0 20–3.0
R
free
(%)
a
29.6 30.3
R
work
(%)
b
27.0 27.0
RMSD
bond
(A
˚
) 0.014 0.010
RMSD
angle
(⬚) 1.47 1.25
Favored 85.6 84.6
Additionally allowed 12.4 13.2
Generously allowed 1.5 1.7
Total atoms (protein) 18,881 18,881
Total atoms (substrates) 190 386
Total atoms (Water) 20 15
a
R
free
is the R value calculated for a test set of reflections, comprising a randomly selected 5% of the data that is not used during refinement.
b
R
work, free
⫽⌺||F
obs
| ⫺ |F
calc
||/|F
obs
|.

Cell
138
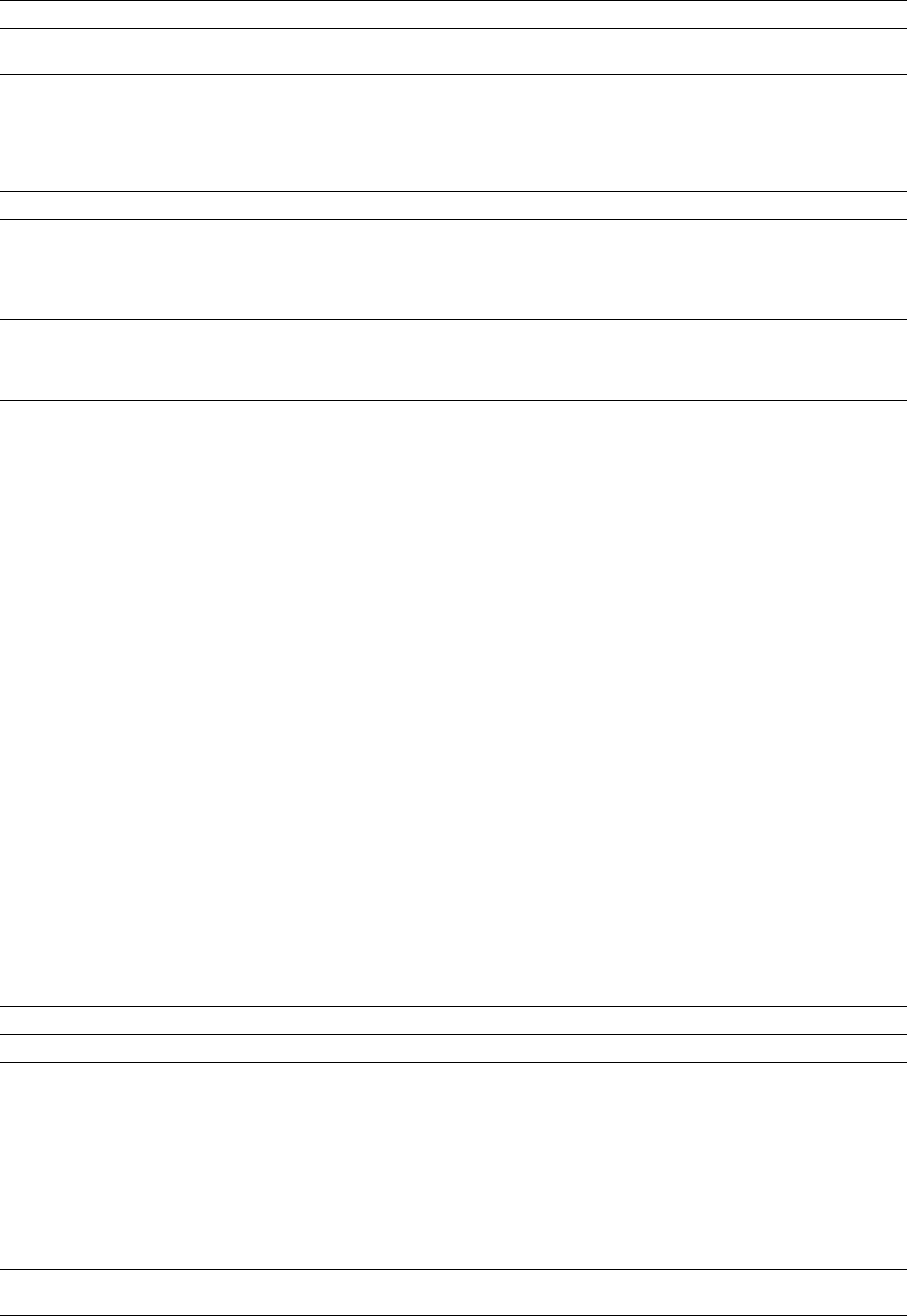
Figure 1. Structure and Topology of the Rho Protomer
(A) Structure of Rho protomer-D showing the relative orientations of the N- (cyan) and C- (red) terminal domains. Each N-terminal domain
consists of a helical bundle and an OB-fold subdomain. The C-terminal domain is connected to the N-terminal domain by an extended linker
(yellow) and consists of a RecA-type ATPase binding fold followed by a small helical subdomain. The P loop (blue), the Q loop (magenta),
and the R loop (green) are highlighted. Secondary structure elements are labeled.
(B) Exploded views of the N- and C-terminal domain interface. Both domains are rotated by 90⬚ opposite to each other. Residues involved
in interdomain contacts are shown as blue rods and labeled.
another by approximately 15⬚ about an axis that lies in ating with mRNA in the absence of accessory factors
(Burgess and Richardson, 2001a, 2001b), this structurala plane perpendicular to the pseudo 6-fold rotation axis
of symmetry (Figure 2C). This swivel generates a helical organization appears consistent with a particle state
competent to load onto extended nucleic acid chains.rise of approximately ⵑ8.5 A
˚
along the rotational axis of
the particle. By propagating these displacements about
the ring of the hexamer, the midpoints of protomers A Rho and Primary Site RNA Interactions
Rho has two distinct nucleic acid binding sites. Theand F at either end of the ring are offset from each other
by 45 A
˚
, giving rise to a 12 A
˚
wide gap that breaches primary mRNA binding sites are formed by the
N-terminal domains, which have the ability to bind eitherthe ring and allows access to the interior of the particle
(Figure 2). It is interesting to note that the width of this single-stranded DNA or RNA (Galluppi and Richardson,
1980; Chen et al., 1986; McSwiggen et al., 1988; Modrakgap is sufficient to allow for the entry of a single-
stranded nucleic acid into the hole of the hexamer. and Richardson, 1994). This feature was used to grow
crystals of Rho in complex with target site mimics com-The intersubunit angular rise present in the Rho com-
plex (15⬚) is within a range seen for the related ATPases posed of either nucleic acid. Earlier studies have shown
that each N-terminal domain binds a dinucleotide seg-RecA and T7 gp4. In RecA, however, the subunit-subunit
rocking angle is more extreme and along the 6
1
helical ment in a network of contacts that explain the preference
of Rho for cytosine (Bogden et al., 1999). The first nucle-axis (18⬚), which leads to filamentous assembly of the
protein (Story et al., 1992). In contrast, for T7 gp4, two otide base packs into a hydrophobic enclosure that is
formed by the side chains of Tyr80, Glu108, and Tyr110,intersubunit rotation angles of 15⬚ in the plane of the
ring are offset by a downward 30⬚ swing every third and is too small to comfortably hold purine bases. For
the second nucleotide, the cytosine base stacks on thesubunit that leads to ring closure (Singleton et al., 2000).
Modulation of subunit-subunit orientations in these pro- aromatic side chain of Phe64, while its O2, N3, and N4
groups interact with the side chains of the neighboringteins is thought to occur in response to the combined
action of ATP turnover and nucleic acid associations, Arg66 and Asp78 (Bogden et al., 1999). No contacts
are seen to the 2⬘ hydroxyl of the bound nucleic acid,which drive either strand exchange (RecA) or DNA trans-
location and unwinding (T7 gp4). In Rho, it appears that explaining why Rho is able to bind both ssDNA and
ssRNA.this common hinging mechanism has been further co-
opted to allow the protein to enter a ring-open conforma- In our full-length Rho complex, the primary RNA bind-
ing sites of the N-terminal domains lie equidistant fromtion without perturbing the oligomerization state of the
protein. Because the interior of Rho is capable of associ- each other about the periphery of the hexamer. There

Rho Termination Factor Structure
139
Figure 2. The Rho Hexamer
(A) Front view of the particle. The six subunits
of Rho pack into an open hexameric ring.
Each protomer is represented by a different
color. The orientation of protomer C (yellow
subunit) is similar to that shown in Figure 1A.
(B) Topdown view of (A). Protomers are la-
beled A–F.
(C) Schematic of the relative rise and offset
of adjacent Rho subunits as they wind about
the pseudo-6-fold axis of the ring (vertical
line). Ovals are colored according to the color
scheme in (A) and (B). The gap between
monomers A and F is 12 A
˚
, and the helical
pitch is 45 A
˚
.
is clear electron density for nucleic acid in the OB-fold density for nucleic acid outside of the primary binding
site.binding cleft of five of the six N-terminal domains. The
one N-terminal domain that lacks density for the DNA Surprisingly, the arrangement between the N- and
C-terminal regions of each protomer is such that theor RNA substrate belongs to protomer B; the helical
bundle of this domain is also disordered. As anticipated, primary RNA binding clefts face inward, toward the cen-
tral hole of the ring (Figure 3A). This finding was unex-each binding site accommodates an oligo-(deoxy)
cytadylic acid dinucleotide as seen for the isolated Rho pected, because earlier data had suggested that the
primary mRNA binding cleft might reside on the outerN-terminal domain/RNA complex solved previously
(Bogden et al., 1999). There is no observable electron periphery of the ring (Yu et al., 2000). Several lines of

Cell
140
Figure 3. Rho RNA Binding Sites
(A) Molecular surface (GRASP [Nicholls et al.,
1991]) of the Rho hexamer. Primary RNA
binding sites in the OB-fold of the N-terminal
domain are colored cyan. Secondary (C-ter-
minal) RNA binding sites in the ATPase do-
main are colored magenta. Nucleic acid
bound at the primary RNA binding sites is
shown as yellow rods. View is the same as in
Figure 2B.
(B) Schematic of the primary (N-terminal) RNA
binding site configuration. The N- and C-ter-
minal domains are colored green and red,
respectively. Solid black lines represent the
positions for the single-stranded nucleic acid,
which binds across the primary RNA binding
site and orients the 3⬘ end toward the hole of
the ring. The broken black line shows the path
needed to be traversed by nucleic acid be-
tween adjacent binding sites.
(C) Rho’s secondary RNA binding site. Stereo
diagram of the Rho hexamer showing the lo-
cation of the P loops (blue), the Q loops (ma-
genta) and the R loops (green). View is from
the “bottom,” rotated 180⬚ from the perspec-
tive of Figure 2B.
evidence, however, indicate that the observed configu- the length of an ssRNA sufficient to span the entire
N-terminal periphery of the ring would be approximatelyration represents the natural architecture of the Rho
particle. First, each of the six protomers in the asymmet- 70–80 bases, a value in excellent agreement with ribo-
nuclease A digestion experiments of RNAs bound to theric unit independently adopts the same conformation
and orientation. Second, there are extensive hydropho- Rho hexamer (Bear et al., 1988; Zhu and von Hippel,
1998b, 1998a).bic contacts between the N- and C-terminal domains,
burying a total surface area of ⵑ850 A
˚
2
per protomer
(Figure 1B). Finally, this arrangement places the ex- Rho and Secondary Site RNA Interactions
mRNA translocation and unwinding are thought to betended linker that connects the N- and C-terminal do-
mains on the exterior of the ring, consistent with prote- catalyzed at Rho’s secondary RNA binding site. This
function depends on two sequence motifs known as thease mapping experiments that have shown this region
to be accessible and labile (Bear et al., 1985; Dolan et Q and R loops (Burgess and Richardson, 2001a; Wei
and Richardson, 2001b, 2001a; Xu et al., 2002). Bothal., 1990).
The relative orientation of the N- and C-terminal do- loops, as predicted from a homology model based on
the F
1
ATPase (Miwa et al., 1995; Burgess and Richard-mains has important consequences for RNA recogni-
tion. The primary RNA binding sites are arranged such son, 2001a), line the interior hole of the hexamer. Each
Q loop lies on the upper segment of the C-terminalthat they bind substrate at an angle of 75⬚ to a plane
perpendicular to the pseudo 6-fold rotation axis of the domain and extends into the center of the ring. The
constellation of the six Q loops in the hexamer togetherring. The substrate is oriented such that the 3⬘ end points
down, toward the interior hole of the ring (Figure 3B). form the narrowest constriction (diameter 20 A
˚
) of the
interior hole (Figures 3A and 3C). Part of the Q loopThis configuration constrains the minimal distance be-
tween the 3⬘ and 5⬘ ends of successive RNA binding of each protomer is disordered, reflecting the intrinsic
mobility of this region, and may result from an absencesites to be 35 A
˚
. As a result, a 12–13 base RNA oligonu-
cleotide would be minimally required to span two adja- of observable contacts with the nucleic acid substrate.
In contrast to the Q loops, the R loops are implicatedcent N-terminal domains and, in contrast to models sug-
gesting that RNA might directly feed from one domain in both ATP and RNA binding (Xu et al., 2002). Each R
loop resides on a segment located at the subunit-sub-to another (Bogden et al., 1999), any RNA long enough
to bridge two or more protomers would be forced to unit interface between the C-terminal domains, and lies
both adjacent to and above the P loop of the ATP bindingzigzag between the primary sites (Figure 3B). In total,

Rho Termination Factor Structure
141
Figure 4. ATP Associations
(A) Stereo view of specific Rho and AMP-PNP interactions. F
o
-F
c
electron density contoured at 2 is shown for the bound nucleotide. Residues
important for nucleotide binding and hydrolysis are shown in ball and stick. Residues from two adjacent C-terminal domains form the active
site and are colored blue and green.
(B) Rho nucleotide binding pocket conformation. The two adjacent C-terminal domains are colored red and gray (protomers D and E). The
active site is partly closed and can bind nucleotide, but does not appear fully organized to carry out hydrolysis. The P loop (blue) is shown
next to the R loop motif (green).
(C) F
1
ATPase nucleotide binding pocket conformation (1bmf). Two ATPase domains are colored yellow and cyan, and bound nucleotide is
shown as black rods. The active site cleft is fully closed and competent to carry out nucleotide hydrolysis.
pocket (Figures 3C and 4B). Part of each R loop also explaining why RNA appears to be readily captured for
translocation following primary site occupancy (Kim andlines the interior hole of the Rho hexamer. The position
of the R loop and its contact with the P loop indicates Patel, 2001).
that this motif could function as part of an allosteric
effector switch that directly couples RNA binding in the Nucleotide Associations and Rho Function
The ATP binding pocket of Rho is located at the interfacehole of the hexamer to the ATP binding and hydrolysis
site (Richardson and Conaway, 1980; Shigesada and between the C-terminal domains of adjacent protomers
(Figure 4B). Part of this region is formed by signatureWu, 1980; Richardson, 1982; Engel and Richardson,
1984; Kim and Patel, 2001). Consistent with this hypoth- sequence motifs such as the adjacent Walker-A
(GXGXXGK(S/T) and Walker-B (D(D/E)XX) segments. Foresis, RNA binding to the secondary state coincides with
closure of the hexameric ring and stimulation of the native crystals grown in the presence of AMPPNP, clear
electron density for bound nucleotide is seen in all sixATPase activity (Gogol et al., 1991; Gan and Richard-
son, 1999), presumably by introducing conformational ATP binding pockets of the hexameric ring (Figure 4A).
In contrast, weak electron density is observed for nucle-changes between subunits and residues around the ATP
binding site (Lowery-Goldhammer and Richardson, otide in the structure determined with the selenomethio-
nine protein, indicating that the occupancy at this site1974). It is interesting to note that the positioning of the
primary RNA binding sites places the 3⬘ RNA end near is low. This difference appears due to the insertion of
the hydrophobic side chain SeMet186 of the seleno-the secondary sites from the outset of mRNA binding,

Cell
142
Figure 5. Schematic Model for Rho Function
Numbers correspond to stages outlined in the
text. Asterisks represent catalytic sites
thought to be competent for ATP hydrolysis.
methionine protein into the adenine binding pocket of ing both an entry point into what would otherwise be
the topologically confined interior of the ring and sug-
each of the six subunits. Nonetheless, superposition
gesting a means by which the protein may enter a nu-
of the ATP bound and -free structures shows that the
cleic acid “loading” configuration. This general organi-
organization and conformation of the ATP binding sites
zation provides an elegant means to ensure that target
are the same.
mRNAs are efficiently captured by the protein, thereby
Cocrystallization (as opposed to soaking) of Rho with
activating the particle and initiating the process of tran-
AMPPNP shows that nucleotide can associate with an
scription termination.
open-ring state of the protein. The interactions between
The significance of the Rho open-ring state may be
Rho and the adenosine base of bound AMPPNP are
applicable to other toroidal helicases and translocases.
typical of those seen for other RecA-type proteins, in
A long-standing puzzle has been to understand how
which aliphatic or aromatic side chains sandwich the
extended nucleic acid segments gain entry to the interior
purine moiety in a “hydrophobic pincer” (Figure 4A)
of such protein rings, where unwinding and transloca-
(Boyer, 1997; Bird et al., 1998a; Singleton et al., 2000;
tion are thought to occur. Many helicases, such as DnaB,
Xu et al., 2000). Other nucleotide associations are not
papilloma virus E1, and the MCM complex, use special-
canonical, however. For example, residues in the
ized loading proteins to accomplish this task (Lusky et
Walker-A and -B regions do not make standard contacts
al., 1993; Seo et al., 1993; Donovan et al., 1997; Tanaka
to the - and ␥-phosphates of AMPPNP, nor is magne-
et al., 1997; Weinreich et al., 1999; Barcena et al., 2001).
sium evident in our structure (Figure 4A). These altered
Others, such as Rho, appear to load onto target sub-
features arise because the subunit-subunit orientation
strates without accessory factors. The ability of Rho to
of the open ring splays apart the ATPase domains as
spontaneously switch between open- and closed-ring
compared to other RecA-type proteins that have formed
structures provides an immediate solution to the loading
competent active sites (Figures 4B and 4C). Given that
problem (Gogol et al., 1991; Yu et al., 2000; Richardson,
the open-ring form of Rho imaged here appears consis-
2002). By utilizing intersubunit hinging motions evident
tent with an mRNA loading state, as opposed to an
in a number of RecA-type ATPases, our structure shows
actively translocating helicase, such atypical nucleotide
how Rho can configure the relative orientation of neigh-
associations are perhaps not surprising, although these
boring RecA-type folds to allow ring opening without
data indicate that ring opening can still occur when Rho
fully exposing subunit interfaces that might otherwise
is bound to ATP, as would be expected when the protein
foster filamentation. As a result, the Rho model may be
operates in vivo.
applicable for understanding how other ring systems
are breached to bind nucleic acids, and may potentially
Rho Architecture and Mechanistic Implications
represent a structural state accessed for some helicases
As a collective, the structures observed here explain
through loader protein activity.
several important features of Rho function. First, Rho’s
Another interesting finding is that the Rho hexamer
primary RNA binding sites line the perimeter of the hex-
can remain open and, thus, competent to bind mRNA
amer and face inward toward the center of the particle.
at its secondary binding sites even when Rho’s ATP and
Second, the primary sites orient the 3⬘ end of nucleic
primary mRNA binding sites are occupied. This observa-
acid target sequences into the interior hole, which is
tion allows us to expand upon models explaining how
formed by motifs known to be critical for coupling mRNA
Rho recognizes rut sites and loads onto RNA in vivo
translocation and unwinding to ATP turnover. Third, the
(Richardson, 2002; Delagoutte and von Hippel, 2003).
Extensive previous work, together with our structuralring of the hexamer is split by a gap ⵑ12 A
˚
wide, provid-
Rho Termination Factor Structure
143
the native protein overexpressed in 2XYT media by induction at an
data, suggest that Rho can oscillate between closed-
OD
600
of 0.3–0.5 with 1 mM IPTG at 37⬚C for 3 hr. Cells were lysed
and open-ring states, either of which may associate with
by sonication in 50 mM Tris-HCl (pH 7.5), 10% glycerol, 50 mM KCl,
ATP and target mRNAs (Figure 5, stages 1 and 2). In the
1 mM TCEP (Tris(2-carboxyethyl)phosphine hydrochloride) on ice.
open state, ATP would be loosely associated with Rho
The protein was twice purified over a POROS-HS column (Perseptive
and only partially liganded as observed here. In contrast,
Biosystems) then further purified with two successive Sephacryl
S-300 (Amersham) sizing columns in 10 mM Tris-HCl (pH 7.5), 50
entry into the closed state would properly position and
mM NaCl, and 1 mM TCEP. The protein at this stage was more than
coordinate nucleotide for hydrolysis. Natural conversion
99% pure as judged by SDS-PAGE and Coomassie staining and
between the two structural states would allow ATP hy-
was monodisperse as judged by dynamic light scattering (DynaPro,
drolysis to be observed in the absence of RNA (Stitt,
model 99-CP, Protein Solutions). Protein was concentrated by ultra-
2001), but turnover would not be fully activated, both
filtration (Centriprep-10, Amicon) to 15 mg/ml. Selenomethionine
because of the lack of RNA effector signals through
protein was overexpressed in minimal media using the protocol of
Van Duyne and coworkers (Van Duyne et al., 1993) and was purified
the Q and R loops and because a population of Rho
similar to native Rho.
molecules might be open at any given moment.
Loading and priming of Rho would begin through the
Crystallization and Data Collection
association of a rut site with the primary RNA binding
Initial crystals of full-length Rho diffracted poorly. Cocrystals with
sites in the N-terminal domains. The configuration of
DNA or RNA substrates resulted in crystals of different morphology
the N-terminal domains automatically orients the 3⬘ end
to those obtained in the absence of nucleic acid and proved instru-
of message toward the interior hole of the protein. Were
mental in solving the Rho structure. A number of single-stranded
the ring to be open during rut site binding, mRNA could
DNA and RNA substrates were tested in cocrystallization trials.
DNAs and RNAs varied in length (6–19 bases) and composition (one
simply thread its way through the gap between mono-
or two binding sites), and were designed based on Rho’s specificity
mers and into the interior of the hexamer to associate
for the unstructured, cytosine-rich mRNA regions present in rut
with the secondary RNA binding sites (Figure 5, stage
sites. The best quality crystals in terms of size, stability, and diffrac-
4). In contrast, if the ring were closed upon rut site
tion were those cocrystallized with a 15-mer DNA substrate (AACC
recognition (Figure 5, stage 3), mRNA entry would need
CAAGAACCCAA) or with an 8-mer RNA substrate (CU)
4
.
to wait until the ring spontaneously opens. It remains a
Cocrystals of Rho in complex with the single-stranded DNA sub-
strate were grown by vapor diffusion at room temperature. Stock
possibility that mRNA binding to the primary, N-terminal
protein solutions were dialyzed in 10 mM Tris-HCl (pH 7.5) and 50
domain binding sites might directly favor ring opening
mM NaCl at 4⬚C prior to crystallization. One volume of protein solu-
(Yu et al., 2000; Kim and Patel, 2001).
tion was mixed with one volume of crystallization solution containing
Once bound in the interior, RNA would be sensed by
100 mM Na•Cacodylate (pH 6.5), 100 mM NaCl, 5% PEG 8K, 40%
the Q and R loop regions, allosterically activating Rho
glycerol, 0.6 mM n-Nonyl--D-thiomaltoside, and 2 mM TCEP; the
and converting the protein into a stable, closed configu-
well solution was diluted with an equivalent amount of H
2
O prior to
sealing the drop over the reservoir. Small crystals appeared over-
ration (Figure 5, stage 5). Based on comparisons with
night and grew to average dimensions of 100 ⫻ 100 ⫻ 200 Min
closed-ring RecA-type proteins, ring closure would pre-
1 week. Crystals were introduced into cryoprotectant, containing a
sumably occur through conformational changes be-
mix of 50 mM Tris-HCl (pH 7.5), 50 mM NaCl, 3% PEG 8K, 25%
tween a subset of the ATPase domains, both forming
glycerol, and 5% PEG 400, in a fast, single step, then flash frozen
competent set of active sites and compensating for the
in liquid nitrogen. The crystals belong to the monoclinic space group
15⬚ subunit-subunit angular rise observed for the open-
C2 with unit cell dimensions of a ⫽ 119.2 A
˚
,b⫽ 205.8 A
˚
,c⫽ 148.0 A
˚
,
and ⫽95.3⬚. There is one hexamer per asymmetric unit.
ring form. Ensuing cycles of ATP turnover would serve to
modulate the relative orientation of neighboring ATPase
domains, providing directed structural changes to pro-
Structure Determination and Refinement
All data were collected at Beamline 8.3.1 at the Advanced Light
pel the protein 5⬘→3⬘ along the message (Brennan et
Source (ALS) using an ADSC Quantum-Q210 CCD detector. Diffrac-
al., 1987; Singleton et al., 2000). As translocation begins,
tion data were processed using DENZO and SCALEPACK (Otwinow-
Rho could either dissociate from the rut site or could
ski, 1997) (Table 1). Initial attempts to calculate phases using seleno-
remain bound to the target motif as predicted by the
methionine MAD or SAD experiments were unsuccessful, so a
tethered tracking model (Figure 5, stages 6 and 7). Al-
tantalum derivative was prepared by soaking a crystal overnight
with saturated solution of Ta
6
Br
12
2
⫹
. Heavy atom positions were
though our structure cannot rule out either mechanism,
identified by SOLVE (Terwilliger and Berendzen, 1999), and
it is interesting to note that the N-terminal domains open
MLPHARE (CCP4, 1994) was used to calculate initial phases to 6.0 A
˚
out from the center of the ring, creating a wide depres-
based on isomorphous and anomalous differences for this derivative
sion across the top of the particle. The size of this de-
(Table 1). Selenium sites (83 out of 96) were located by calculating
pression appears ample enough to remain associated
anomalous difference maps to 3.6 A
˚
resolution using the tantalum
with a RNA segment while single-stranded RNA is
phases (Table 1). The selenium sites were refined initially with
MLPHARE and then with SHARP (de la Fortelle and Bricogne, 1997),
spooled away from the protein during translocation. Fu-
followed by DM solvent flattening (Cowtan, 1994) to yield starting
ture efforts focused toward assembling RNA bound,
electron density maps.
closed ring structures will be needed to help distinguish
Five out of the six N-terminal domains were located in the solvent-
between these models and to better illuminate the physi-
flattened density by the program FFFEAR (Cowtan K., 1998) using
cal coupling between ATP usage, translocation, and un-
the X-ray model 2a8v (Bogden et al., 1999). The sixth domain was
winding.
placed into the map manually and the fit optimized using RSR-RIGID
in O (Jones T.A., 1991). The N-terminal domains were used to derive
the six NCS (noncrystallographic symmetry) operators within theExperimental Procedures
asymmetric unit and the experimental electron density map was
subsequently improved using solvent flattening with NCS multido-Protein Expression and Purification
The open reading frame encoding the full-length Escherichia coli main averaging and phase extension in DM. The C-terminal domain
of the mitochondrial F
1
ATPase model (1bmf) was manually dockedRho protein was PCR amplified and cloned into pET24b. The plasmid
was transformed into the Escherichia coli strain BL21 (pLysS), and and rebuilt into this DM-averaged density using O.
Cell
144
The model was refined with REFMAC5 (Murshudov G.N., 1997), Brennan, C.A., Steinmetz, E.J., Spear, P., and Platt, T. (1990). Speci-
ficity and efficiency of rho-factor helicase activity depends on mag-using ARP (Lamzin and Wilson, 1993) for water building. In the final
cycles of refinement, TLS (translation libration and screw-rotation) nesium concentration and energy coupling to NTP hydrolysis. J.
Biol. Chem. 265, 5440–5447.refinement of rigid groups (Winn et al., 2001) was carried out as
implemented in REFMAC5 (Table 2). The refined selenomethionine
Briercheck, D.M., Wood, T.C., Allison, T.J., Richardson, J.P., and
Rho/DNA structure was used to solve the native Rho/RNA•AMPPNP
Rule, G.S. (1998). The NMR structure of the RNA binding domain of
complex by molecular replacement using the program AMORE (Na-
E. coli rho factor suggests possible RNA-protein interactions. Nat.
vaza, 2001). The resultant solution was DM solvent flattened and
Struct. Biol. 5, 393–399.
multidomain averaged, revealing clear electron density in F
o
-F
c
maps
Brown, S., Brickman, E.R., and Beckwith, J. (1981). Blue ghosts: a
for AMPPNP in all six ATP binding sites. The structure was then
new method for isolating amber mutants defective in essential genes
refined using REFMAC5. The Rho/DNA and the Rho/RNA structures
of Escherichia coli. J. Bacteriol. 146, 422–425.
were refined to a final R
work
27.0% and 27.0% and an R
free
29.6%
Burgess, B.R., and Richardson, J.P. (2001a). RNA passes through
and 30.3%, respectively. A total of 2472 out of 2502 residues are
the hole of the protein hexamer in the complex with the Escherichia
accounted for in either of the two structures (Table 2). Geometric
coli Rho factor. J. Biol. Chem. 276, 4182–4189.
analysis were carried out with Procheck (Laskowski et al., 1993)
and structural superpositions with LSQKAB (CCP4, 1994).
Burgess, B.R., and Richardson, J.P. (2001b). Transcription factor
Rho does not require a free end to act as an RNA-DNA helicase on
Acknowledgments
an RNA. J. Biol. Chem. 276, 17106–17110.
Cavarelli, J., Rees, B., Ruff, M., Thierry, J.C., and Moras, D. (1993).
The authors are grateful to James Holton at Beamline 8.3.1 of the
Yeast tRNA(Asp) recognition by its cognate class II aminoacyl-tRNA
Advanced Light source for assistance with data acquisition and to
synthetase. Nature 362, 181–184.
David King for mass spectroscopy analyses. We would also like to
CCP4 (Collaborative Computational Project 4) (1994). The CCP4
thank James Keck, Deborah Fass, David Akey, Jan Erzberger, and
suite: programs for protein crystallography. Acta Crystallogr. D 50,
Scott Gradia for critical reading the manuscript, as well as members
760–763.
of the Berger Lab for helpful discussions and insights. This work
Chen, C.Y., Galluppi, G.R., and Richardson, J.P. (1986). Transcrip-
was supported by generous assistance from the G. Harold and Leila
tion termination at lambda tR1 is mediated by interaction of rho with
Y. Mathers Charitable Foundation.
specific single-stranded domains near the 3⬘ end of cro mRNA. Cell
46, 1023–1028.
Received: March 19, 2003
Revised: May 20, 2003
Cowtan, K. (1994). A CCP4 density modification package. Joint
Accepted: June 25, 2003
CCP4 ESF-EACBM. Newslett. Prot. Crystallogr. 31, 34–38.
Published: July 10, 2003
Cowtan, K. (1998). Modified phased translation functions and their
application to molecular fragment location. Acta Crystallogr. D 54,
References
750–756.
de la Fortelle, E., and Bricogne, G. (1997). Maximum-likelihood heavy
Abrahams, J.P., Leslie, A.G., Lutter, R., and Walker, J.E. (1994).
atom parameter refinement for multiple isomorphous replacement
Structure at 2.8 A
˚
resolution of F1-ATPase from bovine heart mito-
and multiwavelength anomalous diffraction methods. Methods En-
chondria. Nature 370, 621–628.
zymol. 276, 472–494.
Alifano, P., Rivellini, F., Limauro, D., Bruni, C.B., and Carlomagno,
Delagoutte, E., and von Hippel, P.H. (2003). Helicase mechanisms
M.S. (1991). A consensus motif common to all Rho-dependent pro-
and the coupling of helicases within macromolecular machines. Part
karyotic transcription terminators. Cell 64, 553–563.
II: integration of helicases into cellular processes. Q. Rev. Biophys.
Allison, T.J., Wood, T.C., Briercheck, D.M., Rastinejad, F., Richard-
36, 1–69.
son, J.P., and Rule, G.S. (1998). Crystal structure of the RNA-binding
Dolan, J.W., Marshall, N.F., and Richardson, J.P. (1990). Transcrip-
domain from transcription termination factor rho. Nat. Struct. Biol.
tion termination factor rho has three distinct structural domains. J.
5, 352–356.
Biol. Chem. 265, 5747–5754.
Barcena, M., Ruiz, T., Donate, L.E., Brown, S.E., Dixon, N.E., Rader-
Donovan, S., Harwood, J., Drury, L.S., and Diffley, J.F. (1997).
macher, M., and Carazo, J.M. (2001). The DnaB.DnaC complex: a
Cdc6p-dependent loading of Mcm proteins onto pre-replicative
structure based on dimers assembled around an occluded channel.
chromatin in budding yeast. Proc. Natl. Acad. Sci. USA 94, 5611–
EMBO J. 20, 1462–1468.
5616.
Bear, D.G., Andrews, C.L., Singer, J.D., Morgan, W.D., Grant, R.A.,
Engel, D., and Richardson, J.P. (1984). Conformational alterations
von Hippel, P.H., and Platt, T. (1985). Escherichia coli transcription
of transcription termination protein rho induced by ATP and by RNA.
termination factor rho has a two-domain structure in its activated
Nucleic Acids Res. 12, 7389–7400.
form. Proc. Natl. Acad. Sci. USA 82, 1911–1915.
Galluppi, G.R., and Richardson, J.P. (1980). ATP-induced changes
Bear, D.G., Hicks, P.S., Escudero, K.W., Andrews, C.L., McSwiggen,
in the binding of RNA synthesis termination protein Rho to RNA. J.
J.A., and von Hippel, P.H. (1988). Interactions of Escherichia coli
Mol. Biol. 138, 513–539.
transcription termination factor rho with RNA. II. Electron micros-
copy and nuclease protection experiments. J. Mol. Biol. 199,
Gan, E., and Richardson, J.P. (1999). ATP and other nucleotides
623–635.
stabilize the Rho-mRNA complex. Biochemistry 38, 16882–16888.
Bird, L.E., Brannigan, J.A., Subramanya, H.S., and Wigley, D.B.
Geiselmann, J., Yager, T.D., and von Hippel, P.H. (1992). Functional
(1998a). Characterisation of Bacillus stearothermophilus PcrA heli-
interactions of ligand cofactors with Escherichia coli transcription
case: evidence against an active rolling mechanism. Nucleic Acids
termination factor rho. II. Binding of RNA. Protein Sci. 1, 861–873.
Res. 26, 2686–2693.
Geiselmann, J., Wang, Y., Seifried, S.E., and von Hippel, P.H. (1993).
Bird, L.E., Subramanya, H.S., and Wigley, D.B. (1998b). Helicases:
A physical model for the translocation and helicase activities of
a unifying structural theme? Curr. Opin. Struct. Biol. 8, 14–18.
Escherichia coli transcription termination protein Rho. Proc. Natl.
Acad. Sci. USA 90, 7754–7758.
Bogden, C.E., Fass, D., Bergman, N., Nichols, M.D., and Berger,
J.M. (1999). The structural basis for terminator recognition by the
Gogol, E.P., Seifried, S.E., and von Hippel, P.H. (1991). Structure
Rho transcription termination factor. Mol. Cell 3, 487–493.
and assembly of the Escherichia coli transcription termination factor
rho and its interaction with RNA. I. Cryoelectron microscopic stud-
Boyer, P.D. (1997). The ATP synthase–a splendid molecular ma-
ies. J. Mol. Biol. 221, 1127–1138.
chine. Annu. Rev. Biochem. 66, 717–749.
Brennan, C.A., Dombroski, A.J., and Platt, T. (1987). Transcription Jones, T.A., Zou, J.Y., Cowan, S.W., and Kjeldgaard, M. (1991).
Improved methods for building protein models in electron densitytermination factor rho is an RNA-DNA helicase. Cell 48, 945–952.
Rho Termination Factor Structure
145
maps and the location of errors in these models. Acta Crystallogr. simultaneous interaction at two kinds of nucleic acid-binding sites.
J. Biol. Chem. 257, 5760–5766.A 47, 110–119.
Richardson, J.P., and Conaway, R. (1980). Ribonucleic acid release
Kim, D.E., and Patel, S.S. (2001). The kinetic pathway of RNA binding
activity of transcription termination protein rho is dependent on the
to the Escherichia coli transcription termination factor Rho. J. Biol.
hydrolysis of nucleoside triphosphates. Biochemistry 19, 4293–
Chem. 276, 13902–13910.
4299.
Lamzin, V.S., and Wilson, K.S. (1993). Automated refinement of pro-
Richardson, J.P., and Greenblatt, J. (1996). Escherichia coli and
tein models. Acta Crystallogr. D 49, 129–147.
salmonella. In Cellular and Molecular Biology, Second Edition, F.C.
Laskowski, A.R., MacArthur, W.M., Moss, S.D., and Thornton, M.J.
Neidhardt, R. Curtiss III, J.L. Ingraham, E.C.C. Lin, K.B. Low, B.
(1993). PROCHECK: a program to check the stereochemical quality
Magasanik, W.S. Reznikoff, M. Riley, M. Schaechter, and H.E. Um-
of protein structures. J. Appl. Cryst. 26, 283–291.
barger, eds. (Washington, D.C.: American Society for Microbiology),
Lowery-Goldhammer, C., and Richardson, J.P. (1974). An RNA-
pp. 822–848.
dependent nucleoside triphosphate phosphohydrolase (ATPase)
Richardson, L.V., and Richardson, J.P. (1992). Cytosine nucleoside
associated with rho termination factor. Proc. Natl. Acad. Sci. USA
inhibition of the ATPase of Escherichia coli termination factor rho:
71, 2003–2007.
evidence for a base specific interaction between rho and RNA. Nu-
Lusky, M., Hurwitz, J., and Seo, Y.S. (1993). Cooperative assembly
cleic Acids Res. 20, 5383–5387.
of the bovine papilloma virus E1 and E2 proteins on the replication
Roberts, J.W. (1969). Termination factor for RNA synthesis. Nature
origin requires an intact E2 binding site. J. Biol. Chem. 268, 15795–
224, 1168–1174.
15803.
Sawaya, M.R., Guo, S., Tabor, S., Richardson, C.C., and Ellenberger,
Martinez, A., Burns, C.M., and Richardson, J.P. (1996a). Residues
T. (1999). Crystal structure of the helicase domain from the replica-
in the RNP1-like sequence motif of Rho protein are involved in RNA-
tive helicase-primase of bacteriophage T7. Cell 99, 167–177.
binding affinity and discrimination. J. Mol. Biol. 257, 909–918.
Schindelin, H., Jiang, W., Inouye, M., and Heinemann, U. (1994).
Martinez, A., Opperman, T., and Richardson, J.P. (1996b). Mutational
Crystal structure of CspA, the major cold shock protein of Esche-
analysis and secondary structure model of the RNP1-like sequence
richia coli. Proc. Natl. Acad. Sci. USA 91, 5119–5123.
motif of transcription termination factor Rho. J. Mol. Biol. 257,
Seo, Y.S., Muller, F., Lusky, M., Gibbs, E., Kim, H.Y., Phillips, B.,
895–908.
and Hurwitz, J. (1993). Bovine papilloma virus (BPV)-encoded E2
McSwiggen, J.A., Bear, D.G., and von Hippel, P.H. (1988). Interac-
protein enhances binding of E1 protein to the BPV replication origin.
tions of Escherichia coli transcription termination factor rho with
Proc. Natl. Acad. Sci. USA 90, 2865–2869.
RNA. I. Binding stoichiometries and free energies. J. Mol. Biol. 199,
Shigesada, K., and Wu, C.W. (1980). Studies of RNA release reaction
609–622.
catalyzed by E. coli transcription termination factor rho using iso-
Miwa, Y., Horiguchi, T., and Shigesada, K. (1995). Structural and
lated ternary transcription complexes. Nucleic Acids Res. 8, 3355–
functional dissections of transcription termination factor rho by ran-
3369.
dom mutagenesis. J. Mol. Biol. 254, 815–837.
Singleton, M.R., Sawaya, M.R., Ellenberger, T., and Wigley, D.B.
Modrak, D., and Richardson, J.P. (1994). The RNA-binding domain
(2000). Crystal structure of T7 gene 4 ring helicase indicates a mech-
of transcription termination factor rho: isolation, characterization,
anism for sequential hydrolysis of nucleotides. Cell 101, 589–600.
and determination of sequence limits. Biochemistry 33, 8292–8299.
Steinmetz, E.J., and Platt, T. (1994). Evidence supporting a tethered
Morgan, W.D., Bear, D.G., and von Hippel, P.H. (1983). Rho-depen-
tracking model for helicase activity of Escherichia coli Rho factor.
dent termination of transcription. I. Identification and characteriza-
Proc. Natl. Acad. Sci. USA 91, 1401–1405.
tion of termination sites for transcription from the bacteriophage
Stitt, B.L. (2001). Escherichia coli transcription termination factor
lambda PR promoter. J. Biol. Chem. 258, 9553–9564.
Rho binds and hydrolyzes ATP using a single class of three sites.
Morgan, W.D., Bear, D.G., Litchman, B.L., and von Hippel, P.H.
Biochemistry 40, 2276–2281.
(1985). RNA sequence and secondary structure requirements for
Story, R.M., Weber, I.T., and Steitz, T.A. (1992). The structure of the
rho-dependent transcription termination. Nucleic Acids Res. 13,
E. coli recA protein monomer and polymer. Nature 355, 318–325.
3739–3754.
Sullivan, S.L., and Gottesman, M.E. (1992). Requirement for E. coli
Murshudov G.N., Vagin, A.A., and Dodson, E.J. (1997). Refinement
NusG protein in factor-dependent transcription termination. Cell 68,
of macromolecular structures by the maximum-likelihood method.
989–994.
Acta Crystallogr. D 53, 240–255.
Tanaka, T., Knapp, D., and Nasmyth, K. (1997). Loading of an Mcm
Navaza, J. (2001). Implementation of molecular replacement in
protein onto DNA replication origins is regulated by Cdc6p and
AMoRe. Acta Crystallogr. D Biol. Crystallogr. 57, 1367–1372.
CDKs. Cell 90, 649–660.
Newkirk, K., Feng, W., Jiang, W., Tejero, R., Emerson, S.D., Inouye,
Terwilliger, T.C., and Berendzen, J. (1999). Automated MAD and
M., and Montelione, G.T. (1994). Solution NMR structure of the major
MIR structure solution. Acta Crystallogr. D 55, 849–861.
cold shock protein (CspA) from Escherichia coli: identification of a
Van Duyne, G.D., Standaert, R.F., Karplus, P.A., Schreiber, S.L.,
binding epitope for DNA. Proc. Natl. Acad. Sci. USA 91, 5114–5118.
and Clardy, J. (1993). Atomic structures of the human immunophilin
Nicholls, A., Sharp, K.A., and Honig, B. (1991). Protein folding and
FKBP-12 complexes with FK506 and rapamycin. J. Mol. Biol. 229,
association: insights from the interfacial and thermodynamic prop-
105–124.
erties of hydrocarbons. Proteins 11, 281–296.
von Hippel, P.H., and Delagoutte, E. (2001). A general model for
Oda, T., and Takanami, M. (1972). Observations on the structure of
nucleic acid helicases and their “coupling” within macromolecular
the termination factor rho and its attachment to DNA. J. Mol. Biol.
machines. Cell 104, 177–190.
71, 799–802.
Walstrom, K.M., Dozono, J.M., Robic, S., and von Hippel, P.H. (1997).
Opperman, T., and Richardson, J.P. (1994). Phylogenetic analysis
Kinetics of the RNA-DNA helicase activity of Escherichia coli tran-
of sequences from diverse bacteria with homology to the Esche-
scription termination factor rho. 1. Characterization and analysis of
richia coli rho gene. J. Bacteriol. 176, 5033–5043.
the reaction. Biochemistry 36, 7980–7992.
Otwinowski, Z., and Minor, W. (1997). Processing of X-ray diffraction
Wei, R.R., and Richardson, J.P. (2001a). Identification of an RNA-
data collected in oscillation mode. Methods Enzymol. 276, 307–326.
binding Site in the ATP binding domain of Escherichia coli Rho by
Platt, T. (1994). Rho and RNA: models for recognition and response.
H2O2/Fe-EDTA cleavage protection studies. J. Biol. Chem. 276,
Mol. Microbiol. 11, 983–990.
28380–28387.
Richardson, J. (2002). Rho-dependent termination and ATPases in
Wei, R.R., and Richardson, J.P. (2001b). Mutational changes of con-
transcript termination. Biochim. Biophys. Acta 1577, 251–260.
served residues in the Q-loop region of transcription factor Rho greatly
reduce secondary site RNA-binding. J. Mol. Biol. 314, 1007–1015.Richardson, J.P. (1982). Activation of rho protein ATPase requires
Cell
146
Weinreich, M., Liang, C., and Stillman, B. (1999). The Cdc6p nucleo-
tide-binding motif is required for loading mcm proteins onto chroma-
tin. Proc. Natl. Acad. Sci. USA 96, 441–446.
Winn, M.D., Isupov, M.N., and Murshudov, G.N. (2001). Use of TLS
parameters to model anisotropic displacements in macromolecular
refinement. Acta Crystallogr. D 57, 122–133.
Xu, H., Frank, J., Niedenzu, T., and Saenger, W. (2000). DNA helicase
RepA: cooperative ATPase activity and binding of nucleotides. Bio-
chemistry 39, 12225–12233.
Xu, Y., Kohn, H., and Widger, W.R. (2002). Mutations in the rho
transcription termination factor that affect RNA tracking. J. Biol.
Chem. 277, 30023–30030.
Yu, X., Horiguchi, T., Shigesada, K., and Egelman, E.H. (2000). Three-
dimensional reconstruction of transcription termination factor rho:
orientation of the N-terminal domain and visualization of an RNA-
binding site. J. Mol. Biol. 299, 1279–1287.
Zhu, A.Q., and von Hippel, P.H. (1998a). Rho-dependent termination
within the trp t’ terminator. I. Effects of rho loading and template
sequence. Biochemistry 37, 11202–11214.
Zhu, A.Q., and von Hippel, P.H. (1998b). Rho-dependent termination
within the trp t’ terminator. II. Effects of kinetic competition and rho
processivity. Biochemistry 37, 11215–11222.
Accession Numbers
Coordinates for both Rho complexes have been deposited in the
RCSB PDB database and are available under the accession codes
1PV4 and 1PVO.
